Beyond genes: Protein atlas scores nitrogen fixing duet

The model legume Medicago truncatula. The plant and its bacterial symbiont were used in a landmark University of Wisconsin–Madison study to detail the proteins involved in the process of nitrogen fixation, where plant nutrients are drawn from the atmosphere. Dhileepkumar Jayaraman and Shanmugam Rajasekar/Ane lab
Of the many elusive grails of agricultural biotechnology, the ability to confer nitrogen fixation into non-leguminous plants such as cereals ranks near the very top.
Doing so is a huge challenge because legumes partner with bacteria called rhizobia in a symbiotic waltz that enables plants to draw sustenance from the air and transcend the need for environmentally harmful chemical fertilizers. The natural process is central to the practice of crop rotation, widely used to prevent exhaustion of soil from crops such as corn, which depend on the application of synthetic fertilizers.
The fact that two distinct and very distantly related organisms — a plant and a bacterium — can partner to perform the feat of drawing life-sustaining nitrogen from the atmosphere is just one of the challenges plant engineers face as they seek to confer this quality on other important crops.
The answer to the challenge, however, may be one big step closer with the publication Monday (Oct. 17, 2016) of a massive atlas of plant and bacterial proteins at play as the symbiotic process plays out between plant and microbe.
Writing in the current Nature Biotechnology, a group from the University of Wisconsin–Madison details more than 23,000 plant and bacterial proteins and the molecular controls by which they execute the beneficial relationship. The atlas, possibly the most exhaustive proteomic inventory of any kind to date, shows in minute detail the interplay of proteins as rhizobia colonize root nodules on the model legume Medicago truncatula.
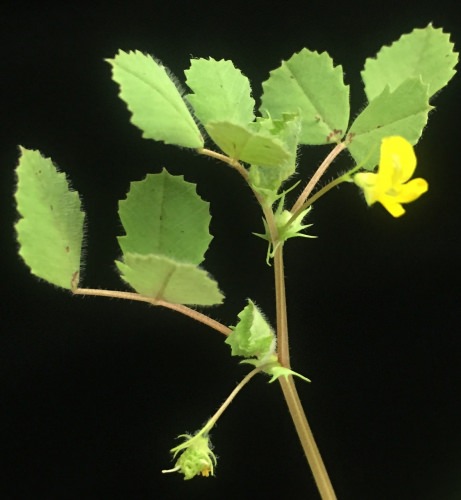

Flowering Medicago truncatula. The plant and its symbiont bacterium were used to detail the thousands of plant and bacterial proteins involved in the process of nitrogen fixation. Dhileepkumar Jayaraman and Shanmugam Rajasekar/Ane lab
“We can see deeper into the proteome than ever before,” explains Joshua Coon, a UW–Madison professor of biomolecular chemistry and chemistry, and a corresponding author of the new atlas. “We’re able to use technology to provide an unprecedented view of these proteins.”
That new picture, he says, takes our understanding of the mechanics of nitrogen fixation to an unprecedented level of detail. Because proteins are regulated by genes, the new atlas could ultimately help inform a strategy for engineering the nitrogen-fixing ability of legumes into other plants.
“Linking the protein information with the genetic networks is important,” notes Jean-Michel Ane, a UW–Madison professor of bacteriology, also a corresponding author of the new report. “It allows us to see patterns by correlating gene expression with proteins.”
The atlas is possibly the most exhaustive proteomic inventory of any kind to date
The new atlas was compiled using potent new mass spectroscopy technology, says Coon, a leading authority on the technique that permits scientists to parse a sample into its many constituent components and measure them in exquisite detail. “The complexity of measuring the number of proteins in a sample is mind-boggling,” Coon says. “Knowing the genes isn’t enough. There are millions and billions of ways proteins can be modified to give them a new mission. All of this information at the protein level is novel, and we can look globally at all these molecules and how they are modified and make some predictions about function.”
The new study, supported mostly by grants from the National Science Foundation, was led by Harald Marx, a postdoctoral fellow in the UW–Madison Genome Center; and Catherine Minogue, a former graduate student in the UW–Madison Department of Chemistry. Michael Sussman, a UW–Madison professor of biochemistry, and Sushmita Roy, a professor of biostatistics and medical informatics, also contributed to the study.
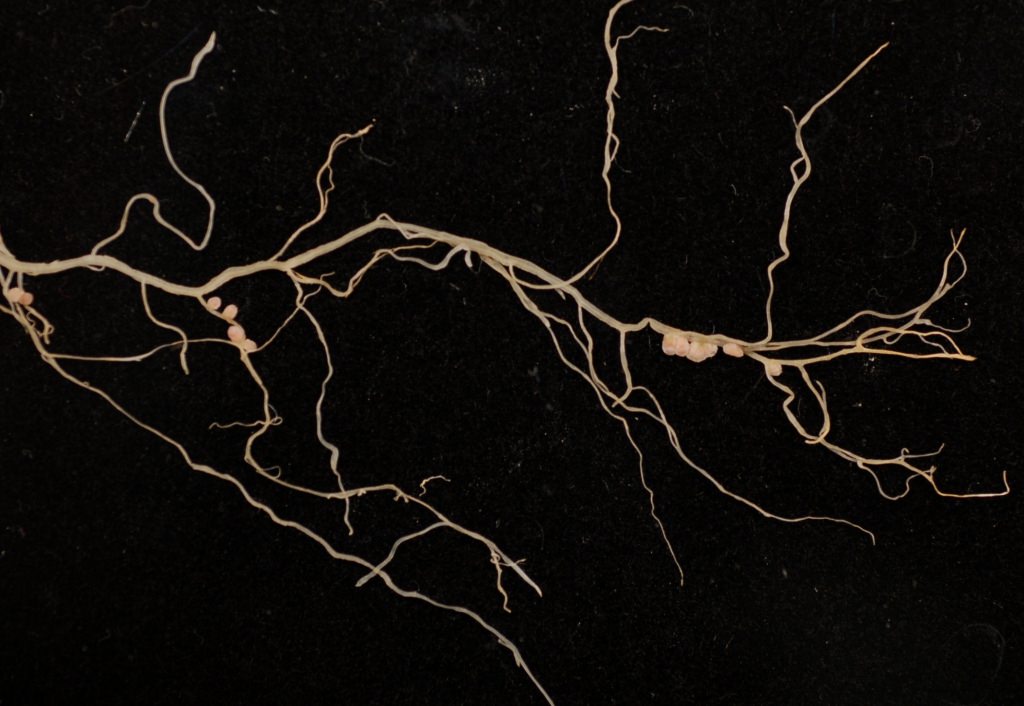
Nodules on the roots of the model legume Medicago truncatula. The root nodules are where the process of nitrogen fixation takes place. Matthew Crook/Ane lab
The Wisconsin researchers stress that while the new protein atlas will be an important cipher for decoding the molecular details of nitrogen fixation symbiosis, the goal of conferring the trait on plants other than legumes remains in the distant future.
The Wisconsin work was conducted using the model legume Medicago truncatula and its rhizobial symbiont Sinorhizobium meliloti, a system developed for genetics research about 20 years ago.
“It is a very close relative to alfalfa,” says Ane, referencing the legume widely used in agriculture as part of crop rotation systems.
Tags: bacteriology, biotechnology, plants




